Recent Progress
The Journal of Biochemistry. Online ahead of print (2024)
Therapeutic strategies targeting cellular senescence for cancer and other diseases.
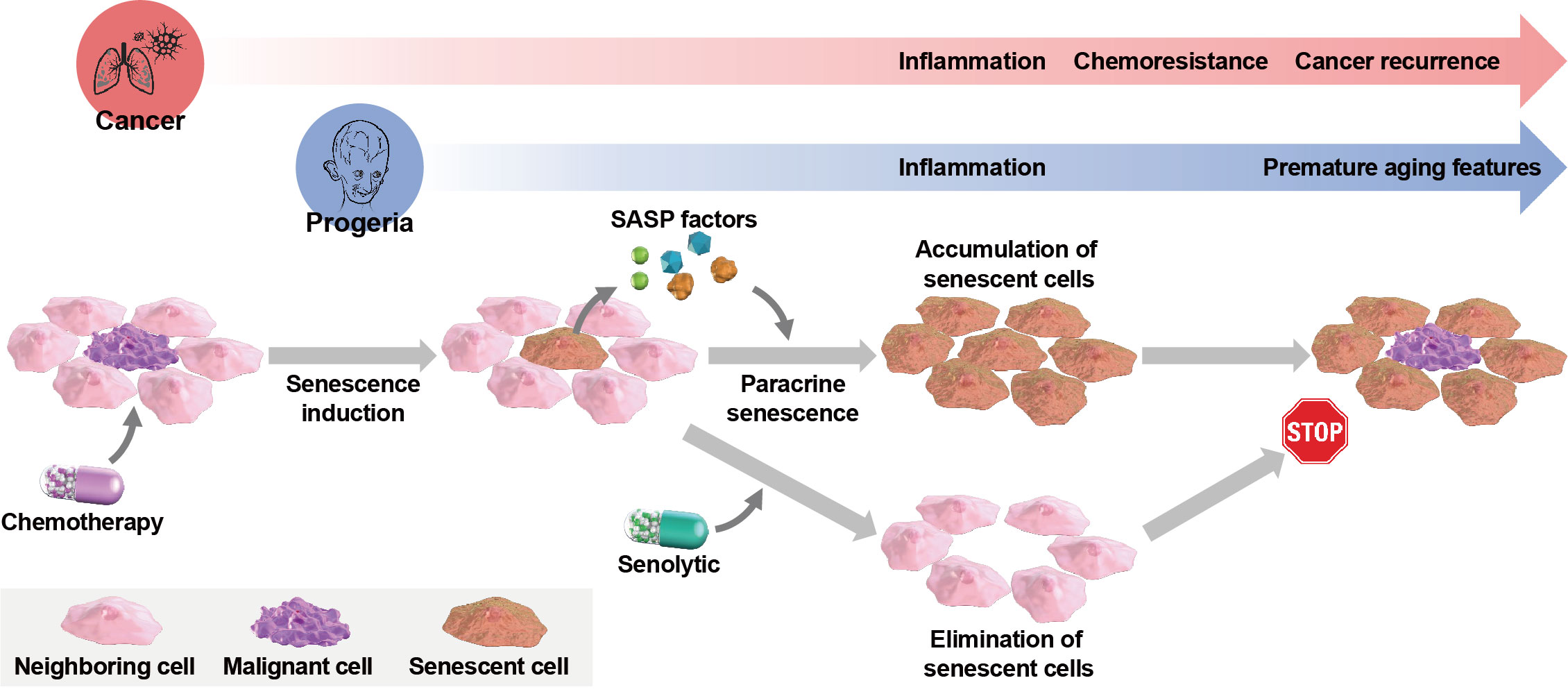
Cellular senescence occurs in response to endogenous or exogenous stresses and is characterized by stable cell cycle arrest, alterations in nuclear morphology, and secretion of proinflammatory factors, referred to as the senescence-associated secretory phenotype (SASP). An increase in senescent cells is associated with the development of several types of cancer and aging-related diseases. Therefore, senolytic agents that selectively remove senescent cells may offer opportunities for developing new therapeutic strategies against such cancers and aging-related diseases. We recently published an article discussing the involvement of senescent cells in certain cancers and aging-related diseases and describing a series of senolytic agents and their utilization in therapeutic strategies.
Nature communications 10, 5688 (2019).
Involvement of condensin in cellular senescence through gene regulation and compartmental reorganization.
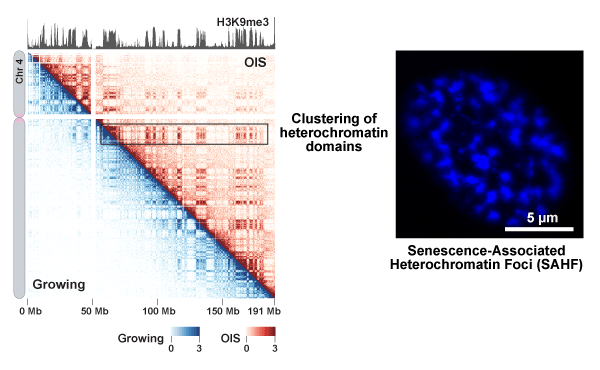
Global genomic reorganization upon cellular senescence
In situ Hi-C experiments were performed using oncogene-induced senescence (OIS) and growing IMR90 cells. Gene contacts for chromosome 4 in OIS (top right) and growing cells (bottom left) are shown at 200 kb resolution. A global H3K9me3 pattern is shown on top. The in situ Hi-C data show that clustering of heterochromatin domains is stronger in OIS cells than proliferating cells. This means that heterochromatin aggregation visualized by DAPI staining (SAHF; right panel) can also be detected by the in situ Hi-C genomics approach.
Nature structural & molecular biology 24, 965–976 (2017).
Architectural alterations of the fission yeast genome during the cell cycle.
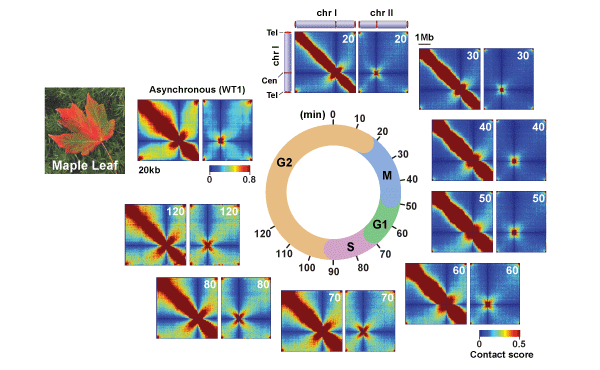
Intra-chromosomal (chr1) and inter-chromosomal (chr1 and chr2) contact maps during the fission yeast cell cycle
Fission yeast cells were cell cycle-synchronized using the cdc25-22 mutant and subjected to in situ Hi-C experiments, and the contact maps were generated throughout the cell cycle. Please note that the global gene contact patterns drastically alter during the cell cycle. The beautiful pattern showing genomic contacts mimics a pattern of maple leaf changing its color during autumn.
Annual Review of Genetics 51, 23-44 (2017)
The Yeast Genomes in Three Dimensions: Mechanisms and Functions
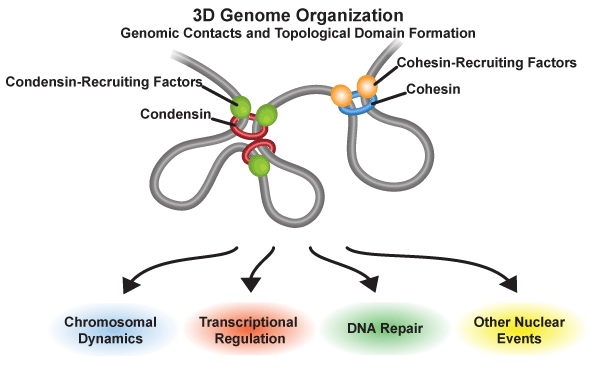
Three-dimensional genome organization and functions in nuclear events
Condensin and cohesin are loaded onto genomic regions by their recruiting factors. They mediate gene contacts and the formation of topological domains. The 3D genome structures organized by condensin and cohesin participate in chromosomal dynamics, transcriptional regulation, DNA repair, and other nuclear events.
Nature Genetics 48, 1242–1252 (2016).
Transcription factors mediate condensin recruitment and global chromosomal organization in fission yeast.
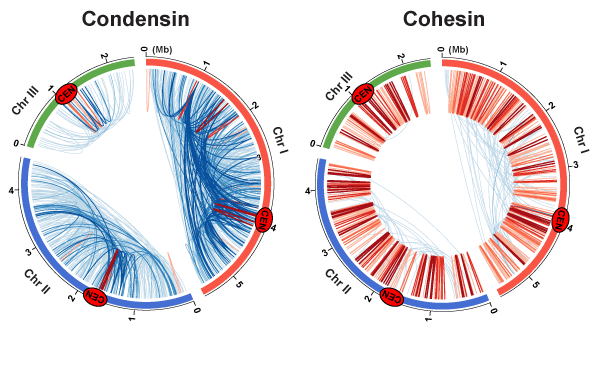
Genome-wide gene contacts mediated by condensin and cohesin
We employed the ChIA-PET genomic method to map condensin (left) and cohesin (right)-mediated gene contacts throughout the fission yeast genome. Blue and red lines represent gene contacts between two loci positioned more and less than 100 kb apart, respectively. Cen indicates centromeres.

